|
ATCC
monoclonal antibody m1 42 3 9 8 Monoclonal Antibody M1 42 3 9 8, supplied by ATCC, used in various techniques. Bioz Stars score: 98/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/monoclonal antibody m1 42 3 9 8/product/ATCC Average 98 stars, based on 1 article reviews
monoclonal antibody m1 42 3 9 8 - by Bioz Stars,
2026-04
98/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp mapk14 mm01301009 m1 Gene Exp Mapk14 Mm01301009 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp mapk14 mm01301009 m1/product/Thermo Fisher Average 93 stars, based on 1 article reviews
gene exp mapk14 mm01301009 m1 - by Bioz Stars,
2026-04
93/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp jak2 hs01078136 m1  Gene Exp Jak2 Hs01078136 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 89/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp jak2 hs01078136 m1/product/Thermo Fisher Average 89 stars, based on 1 article reviews
gene exp jak2 hs01078136 m1 - by Bioz Stars,
2026-04
89/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp cd109 mm00462151 m1  Gene Exp Cd109 Mm00462151 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp cd109 mm00462151 m1/product/Thermo Fisher Average 94 stars, based on 1 article reviews
gene exp cd109 mm00462151 m1 - by Bioz Stars,
2026-04
94/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp gapdh hs99999905 m1  Gene Exp Gapdh Hs99999905 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp gapdh hs99999905 m1/product/Thermo Fisher Average 99 stars, based on 1 article reviews
gene exp gapdh hs99999905 m1 - by Bioz Stars,
2026-04
99/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp ptgs2 mm00478374 m1  Gene Exp Ptgs2 Mm00478374 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp ptgs2 mm00478374 m1/product/Thermo Fisher Average 99 stars, based on 1 article reviews
gene exp ptgs2 mm00478374 m1 - by Bioz Stars,
2026-04
99/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp ddx60 hs01102712 m1  Gene Exp Ddx60 Hs01102712 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp ddx60 hs01102712 m1/product/Thermo Fisher Average 94 stars, based on 1 article reviews
gene exp ddx60 hs01102712 m1 - by Bioz Stars,
2026-04
94/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp pomc mm00435874 m1  Gene Exp Pomc Mm00435874 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp pomc mm00435874 m1/product/Thermo Fisher Average 99 stars, based on 1 article reviews
gene exp pomc mm00435874 m1 - by Bioz Stars,
2026-04
99/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp cdca5 mm01233533 m1  Gene Exp Cdca5 Mm01233533 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 85/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp cdca5 mm01233533 m1/product/Thermo Fisher Average 85 stars, based on 1 article reviews
gene exp cdca5 mm01233533 m1 - by Bioz Stars,
2026-04
85/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp pdgfb hs00966522 m1 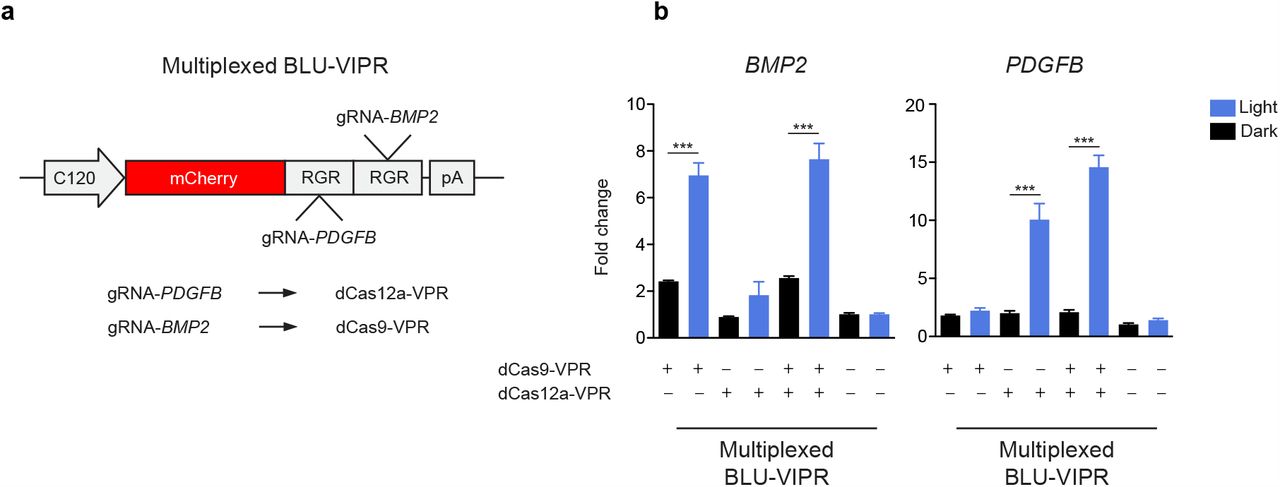 Gene Exp Pdgfb Hs00966522 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 97/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp pdgfb hs00966522 m1/product/Thermo Fisher Average 97 stars, based on 1 article reviews
gene exp pdgfb hs00966522 m1 - by Bioz Stars,
2026-04
97/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp kmt5b hs00992344 m1  Gene Exp Kmt5b Hs00992344 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 91/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp kmt5b hs00992344 m1/product/Thermo Fisher Average 91 stars, based on 1 article reviews
gene exp kmt5b hs00992344 m1 - by Bioz Stars,
2026-04
91/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp upf3b hs00224875 m1  Gene Exp Upf3b Hs00224875 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 86/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp upf3b hs00224875 m1/product/Thermo Fisher Average 86 stars, based on 1 article reviews
gene exp upf3b hs00224875 m1 - by Bioz Stars,
2026-04
86/100 stars
|
Buy from Supplier |
Image Search Results

Journal: bioRxiv
Article Title: The Aryl Hydrocarbon Receptor Controls IFNγ-Induced Immune Checkpoints PD-L1 and IDO via the JAK/STAT Pathway in Lung Adenocarcinoma
doi: 10.1101/2024.08.12.607602
Figure Lengend Snippet: A) CMT167 Ctrl or CMT167 AhR-KO cells were untreated or treated with 100 ng/ml IFNγ. Cd274 mRNA was quantified by RT-qPCR 24h later. Ido1 , Ido2 , Cyp1a1 , and Cyp1b1 mRNA was quantified 72h later. Data are from three independent experiments, each in duplicate or triplicate, are expressed as fold change of Gapdh -normalized means + SE. B) CMT167 Ctrl or CMT167 AhR-KO cells were left untreated or treated for 24h with 100 ng/ml IFNγ and PD-L1 protein expression assayed by western immunoblotting. A representative immunoblot is on the left and β-actin normalized band densities, averaged from three independent experiments, is on the right. (Bands from IFNγ-treated cells reached saturation prior to bands from untreated cells becoming visible). C) CMT167 Ctrl or CMT167 AhR-KO cells were treated for 24h with IFNγ and IDO1 protein expression assayed by western immunoblotting. A representative immunoblot is on the left and β-actin normalized band densities, averaged from three independent experiments, are on the right. D) CMT167 Ctrl or CMT167 AhR-KO cells were treated with 1-1000 ng/ml IFNγ for 24h and Kyn released into the media quantified via colorimetric assay. Data are averaged from two independent experiments each in quadruplicate + SE. E) CMT167 Ctrl or CMT167 AhR-KO cells were treated with 100 ng/ml IFNγ and Jak2 and Stat1 mRNA quantified 24h later. RT-qPCR data are from three independent experiments, each in triplicate, and presented as Gapdh -normalized means + SE. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001 (Student’s t-test, equal variance).
Article Snippet: The following TaqMan assays were purchased from
Techniques: Quantitative RT-PCR, Expressing, Western Blot, Colorimetric Assay

Journal: bioRxiv
Article Title: The Aryl Hydrocarbon Receptor Controls IFNγ-Induced Immune Checkpoints PD-L1 and IDO via the JAK/STAT Pathway in Lung Adenocarcinoma
doi: 10.1101/2024.08.12.607602
Figure Lengend Snippet: A) A549 Ctrl or A549 AhR-KO cells were untreated or treated with 100 ng/ml IFNγ for 24h and CD274 , IDO1, and CYP1B1 expression quantified by RT-qPCR. Data from four experiments, each in triplicate, are represented as fold change of GAPDH -normalized means + SE. B) The percent positive PD-L1 + cells treated as in ( A ) was quantified by flow cytometry. Data from three experiments, each in triplicate, are presented as mean percent PD-L1 + + SE. C) The baseline percent of Kyn + A549 ctrl and A549 AhR- KO cells in two experiments, each in triplicate, was determined by flow cytometry. D) A549 Ctrl or A549 AhR-KO cells were treated with 0-1000 ng/ml IFNγ for 24h and Kyn release quantified by the Kyn-specific colorimetric assay using a standard Kyn curve. Data from two experiments, each in quadruplicate, are presented as average μM Kyn + SE. E) A549 Ctrl or A549 AhR-KO cells were left untreated or treated with 100 ng/ml IFNγ for 24h and baseline or 100 ng/ml IFNγ-induced JAK2 , STAT1, and STAT3 expression quantified by RT-qPCR. Data from four experiments, each in triplicate, are presented as average fold change of Gapdh -normalized means + SE. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001 (Student’s t-test, equal variance).
Article Snippet: The following TaqMan assays were purchased from
Techniques: Expressing, Quantitative RT-PCR, Flow Cytometry, Colorimetric Assay

Journal: bioRxiv
Article Title: The Aryl Hydrocarbon Receptor Controls IFNγ-Induced Immune Checkpoints PD-L1 and IDO via the JAK/STAT Pathway in Lung Adenocarcinoma
doi: 10.1101/2024.08.12.607602
Figure Lengend Snippet: A) Expression of Cd274 , Ido1 , Ido2 , Cyp1a1 , and Cyp1b1 mRNA was quantified by RT-qPCR in control and CMT167 AhR-KO cells (left two bars in each plot) or after 72 hours of treatment with 10 µM benzo(a)pyrene (B(a)P)(right two bars in each plot). Data from three independent experiments, each in duplicate or triplicate are presented as Gapdh -normalized means + SE. B) Protein extracted from cells treated as in ( A ) was probed by western immunoblotting for PD-L1 and, as a loading control, β-actin. One of three representative western blots is shown on the left and β-actin-normalized protein band densities are on the right. Band density data are presented as means from three experiments, each in triplicate, + SE. C) The percentage of Kyn + CMT167 WT or CMT167 AhR-KO cells treated with vehicle or B(a)P as in ( A ) was quantified by flow cytometry. Data from two experiments, each in triplicate, are presented as means + SE from. *p<0.05, **p<0.01, ****p<0.0001 (Student’s t-test, equal variance).
Article Snippet: The following TaqMan assays were purchased from
Techniques: Expressing, Quantitative RT-PCR, Control, Western Blot, Flow Cytometry

Journal: bioRxiv
Article Title: The Aryl Hydrocarbon Receptor Controls IFNγ-Induced Immune Checkpoints PD-L1 and IDO via the JAK/STAT Pathway in Lung Adenocarcinoma
doi: 10.1101/2024.08.12.607602
Figure Lengend Snippet: A) CMT167 Ctrl or CMT167 AhR-KO cells were untreated or treated with 100 ng/ml IFNγ. Cd274 mRNA was quantified by RT-qPCR 24h later. Ido1 , Ido2 , Cyp1a1 , and Cyp1b1 mRNA was quantified 72h later. Data are from three independent experiments, each in duplicate or triplicate, are expressed as fold change of Gapdh -normalized means + SE. B) CMT167 Ctrl or CMT167 AhR-KO cells were left untreated or treated for 24h with 100 ng/ml IFNγ and PD-L1 protein expression assayed by western immunoblotting. A representative immunoblot is on the left and β-actin normalized band densities, averaged from three independent experiments, is on the right. (Bands from IFNγ-treated cells reached saturation prior to bands from untreated cells becoming visible). C) CMT167 Ctrl or CMT167 AhR-KO cells were treated for 24h with IFNγ and IDO1 protein expression assayed by western immunoblotting. A representative immunoblot is on the left and β-actin normalized band densities, averaged from three independent experiments, are on the right. D) CMT167 Ctrl or CMT167 AhR-KO cells were treated with 1-1000 ng/ml IFNγ for 24h and Kyn released into the media quantified via colorimetric assay. Data are averaged from two independent experiments each in quadruplicate + SE. E) CMT167 Ctrl or CMT167 AhR-KO cells were treated with 100 ng/ml IFNγ and Jak2 and Stat1 mRNA quantified 24h later. RT-qPCR data are from three independent experiments, each in triplicate, and presented as Gapdh -normalized means + SE. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001 (Student’s t-test, equal variance).
Article Snippet: The following TaqMan assays were purchased from
Techniques: Quantitative RT-PCR, Expressing, Western Blot, Colorimetric Assay

Journal: bioRxiv
Article Title: The Aryl Hydrocarbon Receptor Controls IFNγ-Induced Immune Checkpoints PD-L1 and IDO via the JAK/STAT Pathway in Lung Adenocarcinoma
doi: 10.1101/2024.08.12.607602
Figure Lengend Snippet: A) A549 Ctrl or A549 AhR-KO cells were untreated or treated with 100 ng/ml IFNγ for 24h and CD274 , IDO1, and CYP1B1 expression quantified by RT-qPCR. Data from four experiments, each in triplicate, are represented as fold change of GAPDH -normalized means + SE. B) The percent positive PD-L1 + cells treated as in ( A ) was quantified by flow cytometry. Data from three experiments, each in triplicate, are presented as mean percent PD-L1 + + SE. C) The baseline percent of Kyn + A549 ctrl and A549 AhR- KO cells in two experiments, each in triplicate, was determined by flow cytometry. D) A549 Ctrl or A549 AhR-KO cells were treated with 0-1000 ng/ml IFNγ for 24h and Kyn release quantified by the Kyn-specific colorimetric assay using a standard Kyn curve. Data from two experiments, each in quadruplicate, are presented as average μM Kyn + SE. E) A549 Ctrl or A549 AhR-KO cells were left untreated or treated with 100 ng/ml IFNγ for 24h and baseline or 100 ng/ml IFNγ-induced JAK2 , STAT1, and STAT3 expression quantified by RT-qPCR. Data from four experiments, each in triplicate, are presented as average fold change of Gapdh -normalized means + SE. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001 (Student’s t-test, equal variance).
Article Snippet: The following TaqMan assays were purchased from
Techniques: Expressing, Quantitative RT-PCR, Flow Cytometry, Colorimetric Assay

Journal: bioRxiv
Article Title: Concomitant suppression of COX-1 and COX-2 is insufficient to induce enteropathy associated with chronic NSAID use
doi: 10.1101/2024.11.22.624882
Figure Lengend Snippet: ( A ) Gene expression via qPCR for Ptgs1 and Ptgs2 , which encode COX-1 and COX-2, respectively, in both lung and small intestine tissue. **p<0.01, ***p<0.001, ****p<0.0001 by unpaired t test.
Article Snippet: The following TaqMan primers were used: Ptgs1/Cox-1 : Mm00477214_m1 (Life Tech / Invitrogen / ABI, Carlsbad, CA) Ptgs2/Cox-2 :
Techniques: Expressing

Journal: bioRxiv
Article Title: Intrinsic OASL expression licenses interferon induction during influenza A virus infection
doi: 10.1101/2025.03.14.643375
Figure Lengend Snippet: A ) UMAP visualization of scRNA-seq data from HBECs, with cells labeled based on cell types. B ) Proportion of cells expressing OASL, IFIT3, DDX60, ISG15, or IRF1 and C) expression counts divided by donor. D) The cell type composition of positive cells for each candidate gene separated and colored by cell types. E) Fraction of cells within each cell type expressing the candidate ISGs.
Article Snippet: Reactions were incubated at 45°C for 50 min and 95°C for 2 min and held at 4°C; cDNA was stored at -20 and subsequently used for quantitative PCR. qPCR reactions were setup by combining 10 μL of TaqMan fast advanced master mix (Applied Biosystems), 1 μL of probe ( OASL; Applied Biosystems: Hs00984387_m1, IFIT3 ; Applied Biosystems: Hs01922752_s1, DDX60 ; Applied Biosystems:
Techniques: Labeling, Expressing

Journal: bioRxiv
Article Title: Light induced expression of gRNA allows for optogenetic gene editing of T lymphocytes in vivo
doi: 10.1101/2023.11.09.566272
Figure Lengend Snippet: a , Ribozyme-gRNA-ribozyme (RGR) design for multiplexed activation of dCas12a-VPR and dCas9-VPR. b , Multiplexed, orthogonal optogenetic dCas12a-VPR and Cas9-VPR to activate the endogenous genes PDGFB and BMP2 . HEK293T cells were transfected with BLU-VIPR-multiplexed, and either dCas12a-VPR, dCas9-VPR, or both. PDGFB and BMP2 expression was assayed after 48 hours of light stimulation, normalized to HPRT1 expression and fold change is relative to untransfected samples kept under dark conditions. The experiment was performed with n=5. One-way ANOVA with Tukey post hoc test was used to identify statistical significance (***p<0.001).
Article Snippet: For gene expression analysis we performed RT-qPCR using TaqManTM Universal Master Mix II, no UNG (
Techniques: Activation Assay, Transfection, Expressing